Finnley Howald
Visual computing at the intersection of real-time graphics, machine learning & computational biology
WORK EXPERIENCE
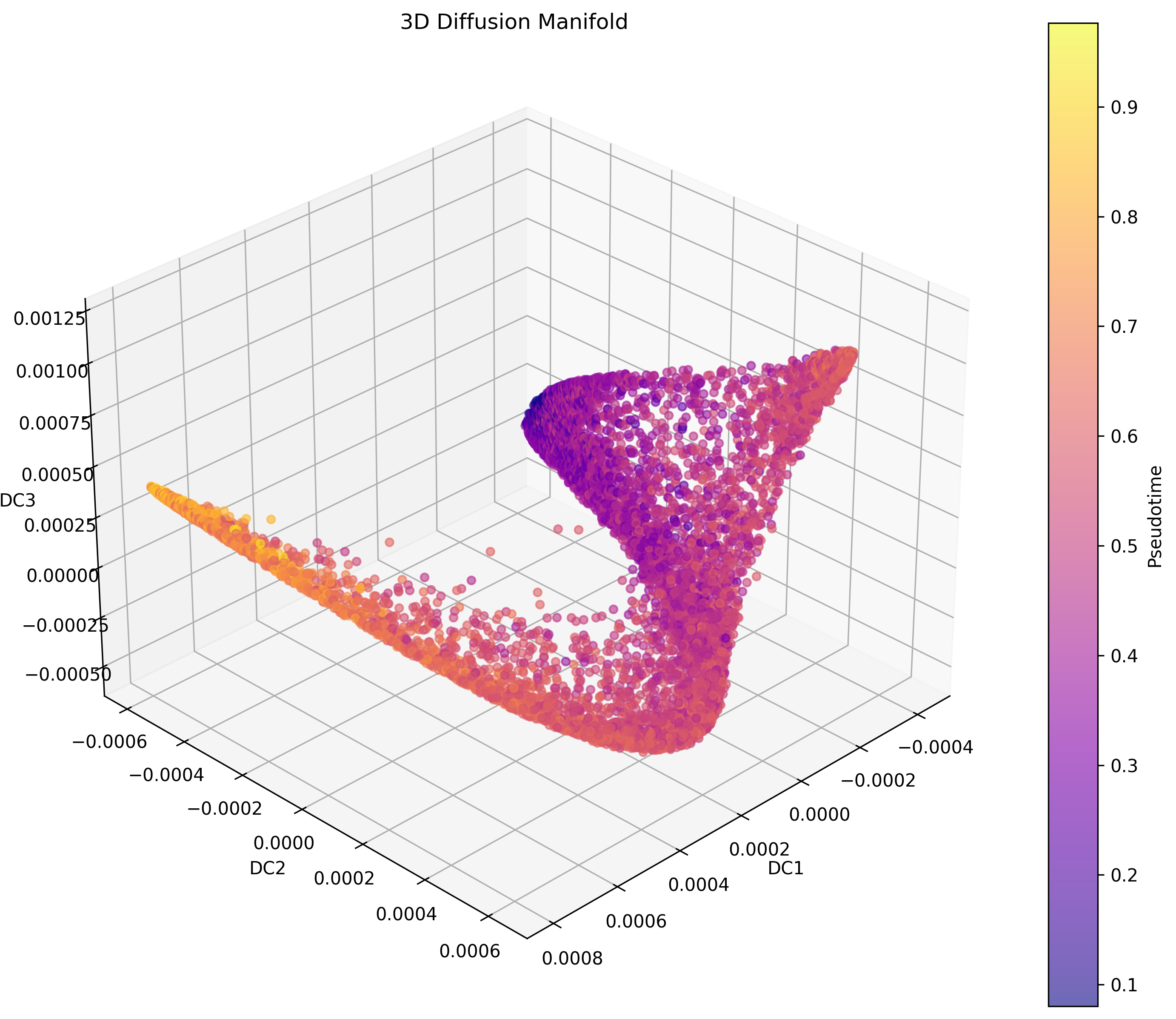
Manifold visualization of morphological trajectories across high-dimensional biological state space.
Built a machine learning pipeline to map cancer progression trajectories from whole-slide histopathology images. Combines deep learning tissue embeddings (Phikon) with diffusion pseudotime analysis to construct a spatial reference atlas showing how breast cancer morphology evolves within and across patient samples.
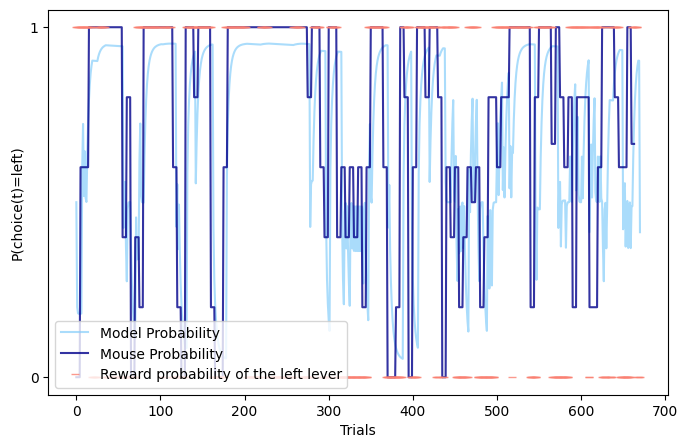
RL simulation traces and choice dynamics used to benchmark learning algorithms against synthetic behaviour.
Implemented and benchmarked RL models (Q-Learning, Double Q, Choice Kernel, Forgetting) for probabilistic choice in mice, with a Python pipeline for cleaning, simulation, and synthetic datasets, plus hyperparameter tuning to improve fit quality and parameter recovery (Double Q + Forgetting strongest).
PROJECTS
PRJ_01
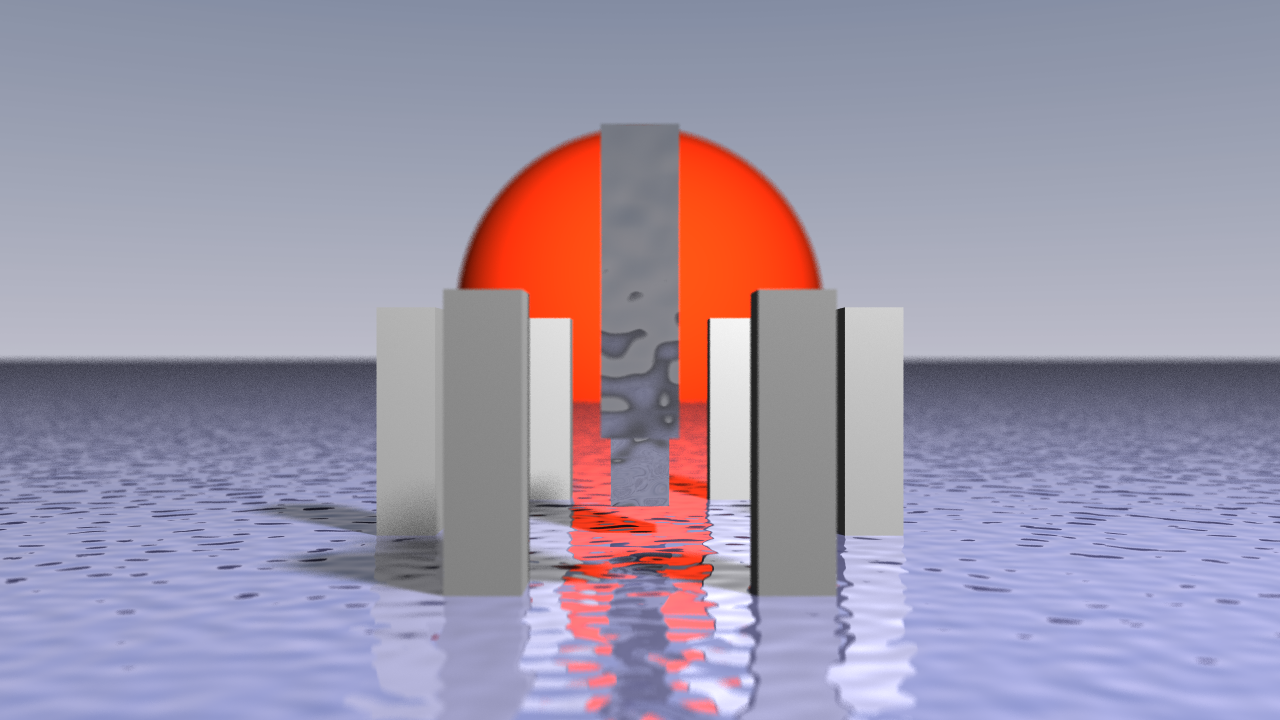

Ray Tracer using Taichi
A ray tracer built using the Taichi programming language, rendering physically accurate reflections, shadows and lighting across complex 3D scenes.
PRJ_02
Mesh Simplification using Quadratic Error
Implementation of the quadratic error metrics algorithm for progressive mesh simplification, preserving visual fidelity while drastically reducing polygon count.
PRJ_03
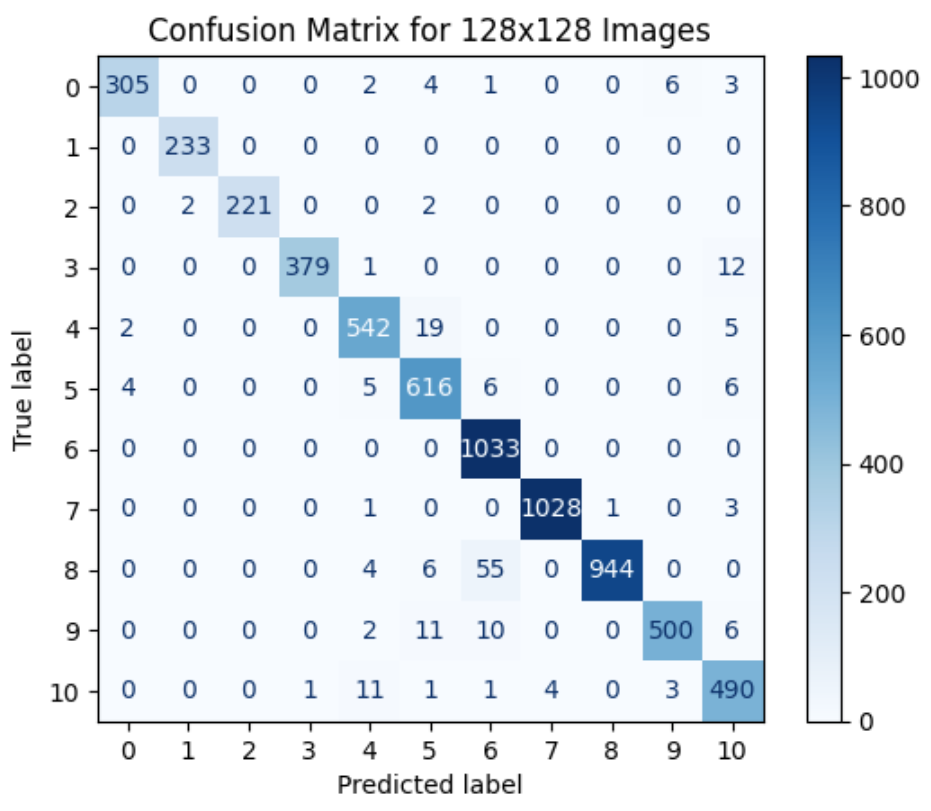
Benchmarking MLPs vs CNNs on Medical Images
Benchmarking scratch-built multilayer perceptrons against convolutional neural networks on medical image classification tasks, evaluating accuracy, convergence and generalization.
PRJ_04

Biological Breast Cancer Simulation
Developed biological models for breast cancer using Python and Compucell3D. Analyzed simulation results to aid in cancer research and shared findings with lab members.
GET IN TOUCH
finnley.howald@gmail.com Master's of Visual Computing @ Simon Fraser University & B.Sc of Computer Science @ McGill University. Currently accepting inquiries for collaborative research, internships, and opportunities at the intersection of computer graphics, machine learning and biological simulation.